By Dmytro Fishman, Junior Lecturer of Data Science

The human organism is a profoundly complex mechanism, consisting of trillions of cells, each powered by about 42 million protein molecules that interact with one another in a gazillion of ways. Recently, scientists developed technologies (i.e. high-throughput screening) that enable us to listen and record data from many of these interactions. Aside from being useful for understanding human biology, this data is utterly difficult to analyse. It is simply too large to use conventional tools such as MS Excel. This is where automated methods such as Machine Learning algorithms come in handy.
Due to their inherent ability to autonomously learn interesting patterns from vast amounts of data, machine learning models have become very popular in biomedical studies. Hence, the number of publications that use these methods has risen significantly over the past few decades.
However, building machine learning models that work in practice has been associated with a number of implicit requirements, such as for example validating models’ predictive power by measuring it on unseen data. At present only a fraction of published papers adheres to such standards, putting the research in jeopardy.
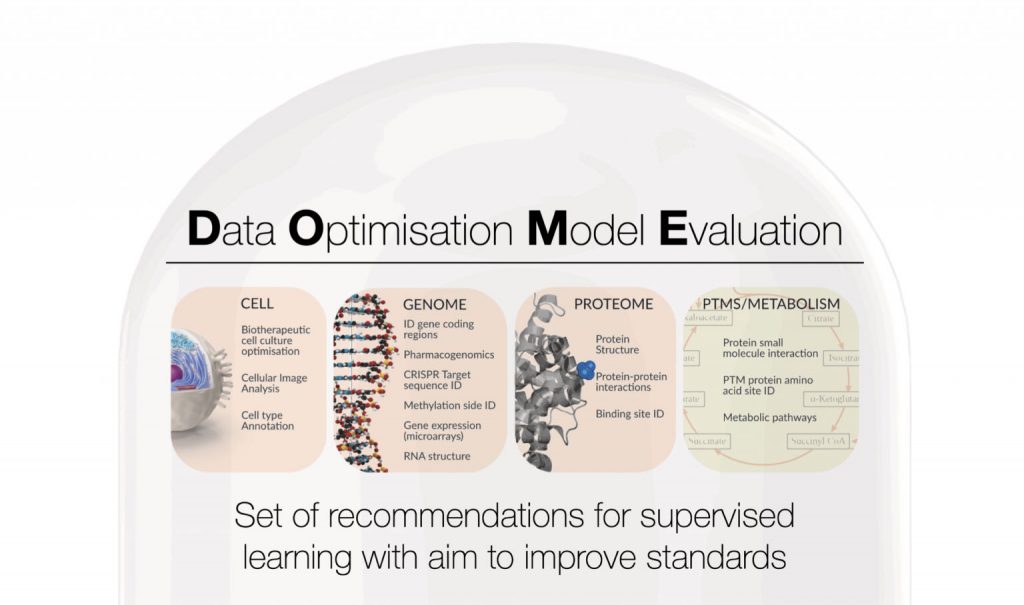
To address this growing gap between the number of publications and the lack of coherence I together with my co-authors from the ELIXIR Machine Learning Focus Group propose a set of community-guided recommendations on how to build machine learning models. More specifically, we have designed minimal requirements asked in a form of questions to machine learning implementers that if followed can ensure the correctness of results.
| Broad topic | Be on the lookout for | Consequences | Recommendation(s) |
| Data | Data size & quality Appropriate partitioning, dependence between train and test data. Class imbalance No access to data | Data not representative of domain application. Unreliable or biased performance evaluation. Cannot check data credibility. | Independence of optimization (training) and evaluation (testing) sets. (requirement) This is especially important for meta algorithms, where the independence of multiple training sets must be shown to be independent of the evaluation (testing) sets. Release data preferably using appropriate long-term repositories, including exact splits (requirement) Sufficient evidence of data size & distribution is representative of the domain. (recommendation) |
| Optimization | Overfitting, underfitting and illegal parameter tuning Imprecise parameters and protocols given. | Reported performance too optimistic or too pessimistic. Models noise or miss relevant relationships. Results are not reproducible. | Clear statement that evaluation sets were not used for feature selection, pre-processing steps or parameter tuning. (requirement) Reporting indicators on training and testing data that can aid in assessing the possibility of under/overfitting e.g. train vs. test error. (requirement) Release definitions of all algorithmic hyper-parameters, regularization protocols, parameters and optimization protocol. (requirement) For neural networks, release definitions of train and learning curves. (recommendation) Include explicit model validation techniques, such as N-fold Cross validation. (recommendation) |
| Model | Unclear if black box or interpretable model No access to: resulting source code, trained models & data Execution time is impractical | An interpretable model shows no explainable behaviour Cannot cross compare methods, reproducibility, & check data credibility. Model takes too much time to produce results | Describe the choice of black box / interpretable model. If interpretable show examples of it doing so. (requirement). Release of: documented source code + models + executable + UI/webserver + software containers. (recommendation) Report execution time averaged across many repeats. If computationally tough compare to similar methods (recommendation) |
| Evaluation | Performance measures inadequate No comparisons to baselines or other methods Highly variable performance. | Biased performance measures reported. The method is falsely claimed as state-of-the-art. Unpredictable performance in production. | Compare with public methods & simple models (baselines). (requirement) Adoption of community validated measures and benchmark datasets for evaluation. (requirement) Comparison of related methods and alternatives on the same dataset. (recommendation) Evaluate performance on a final independent hold-out set. (recommendation) Confidence intervals/error intervals and statistical tests to gauge prediction robustness. (requirement) |
We focused on four key components of any machine learning algorithm: data, optimisation, model, and evaluation (DOME). The main objective of this effort is to increase the reproducibility and clarity of ML methods published in the field of biology.
DOME recommendations were developed through the ELIXIR Machine Learning Focus Group. ELIXIR, initially established in 2014, is now a mature intergovernmental European infrastructure for biological data and represents over 220 research organizations in 22 countries across many aspects of bioinformatics. Over 700 national experts participate in the development and operation of national services that contribute to data access, integration, training, and analysis for the research community. Over 50 of these experts involved in the field of ML have established the ELIXIR Machine Learning Focus Group, which holds meetings to develop and refine recommendations based on a broad consensus.
DOME recommendations have recently been published in Nature Methods.